 Your new post is loading...
 Your new post is loading...
How to use this site to your advantage ... and not get lost
How to benefit the most of this site?
Just follow the steps as below:
- The first possibility (and a highly recommended one) is just to visit it frequently, in order to stay aware of the newly published articles or sources of information as soon as they are posted.
- Further, since the most recently 3,760 postings from the 7,320 originally posted are presently available (as of October 10, 2023) which are arranged as per date of posting, you can do a search according to your specific interests. In doing that, you go to the upper right corner ("Search in topics" depicted with a label), where you can just use the descriptors that are available there; i.e. "Tags" which are ordered alphabetically.). Another possibility is to type there a keyword (or an entire phrase) that can be the name of an author or a word/phrase contained in the title/abstract or anything you deem relevant.
- That way you will be shown a reduced number of sources being more relevant to your specific interest(s).
Hoping this will be useful and waiting for feedback to keep improving the site, I wish all the best
Julio Retamales (the curator)
NOTE: Certainly, given the sheer number of articles being published currently on the relevant issues, no claim for completeness can be provided. Therefore, only samples of papers and/or sources arbitrarily selected by the curator are posted here, intending to show the diversity of phenomena in which plant hormones can be involved.
Authors: Ishfaq Majid Hurrah, Amit Kumar and Nazia Abbas.
Planta (2024)
Main conclusion: Overexpression of Artemisia annua jasmonic acid carboxyl methyltransferase (AaJMT) leads to enhanced artemisinin content in Artemisia annua.
Abstract: "Artemisinin-based combination therapies remain the sole deterrent against deadly disease malaria and Artemisia annua remains the only natural producer of artemisinin. In this study, the 1101 bp gene S-adenosyl-l-methionine (SAM): Artemisia annua jasmonic acid carboxyl methyltransferase (AaJMT), was characterised from A. annua, which converts jasmonic acid (JA) to methyl jasmonate (MeJA). From phylogenetic analysis, we confirmed that AaJMT shares a common ancestor with Arabidopsis thaliana, Eutrema japonica and has a close homology with JMT of Camellia sinensis. Further, the Clustal Omega depicted that the conserved motif I, motif III and motif SSSS (serine) required to bind SAM and JA, respectively, are present in AaJMT. The relative expression of AaJMT was induced by wounding, MeJA and salicylic acid (SA) treatments. Additionally, we found that the recombinant AaJMT protein catalyses the synthesis of MeJA from JA with a Km value of 37.16 µM. Moreover, site-directed mutagenesis of serine-151 in motif SSSS to tyrosine, asparagine-10 to threonine and glutamine-25 to histidine abolished the enzyme activity of AaJMT, thus indicating their determining role in JA substrate binding. The GC–MS analysis validated that mutant proteins of AaJMT were unable to convert JA into MeJA. Finally, the artemisinin biosynthetic and trichome developmental genes were upregulated in AaJMT overexpression transgenic lines, which in turn increased the artemisinin content."
Authors: J. S. Chen, S. T. Wang, Q. Mei, T. Sun, J. T. Hu, G. S. Xiao, H. Chen and Y. H. Xuan.
Plant Molecular Biology (2024)
Abstract: "Plants have a variety of regulatory mechanisms to perceive, transduce, and respond to biotic and abiotic stress. One such mechanism is the calcium-sensing CBL–CIPK system responsible for the sensing of specific stressors, such as drought or pathogens. CBLs perceive and bind Calcium (Ca2+) in response to stress and then interact with CIPKs to form an activated complex. This leads to the phosphorylation of downstream targets, including transporters and ion channels, and modulates transcription factor levels and the consequent levels of stress-associated genes. This review describes the mechanisms underlying the response of the CBL–CIPK pathway to biotic and abiotic stresses, including regulating ion transport channels, coordinating plant hormone signal transduction, and pathways related to ROS signaling. Investigation of the function of the CBL–CIPK pathway is important for understanding plant stress tolerance and provides a promising avenue for molecular breeding."
Author: Peter V. Minorsky.
Plant Signaling & Behavior (2024)
Abstract: "The 21st-century “plant neurobiology” movement is an amalgam of scholars interested in how “neural processes”, broadly defined, lead to changes in plant behavior. Integral to the movement (now called plant behavioral biology) is a triad of historically marginalized subdisciplines, namely plant ethology, whole plant electrophysiology and plant comparative psychology, that set plant neurobiology apart from the mainstream. A central tenet held by these “triad disciplines” is that plants are exquisitely sensitive to environmental perturbations and that destructive experimental manipulations rapidly and profoundly affect plant function. Since destructive measurements have been the norm in plant physiology, much of our “textbook knowledge” concerning plant physiology is unrelated to normal plant function. As such, scientists in the triad disciplines favor a more natural and holistic approach toward understanding plant function. By examining the history, philosophy, sociology and psychology of the triad disciplines, this paper refutes in eight ways the criticism that plant neurobiology presents nothing new, and that the topics of plant neurobiology fall squarely under the purview of mainstream plant physiology. It is argued that although the triad disciplines and mainstream plant physiology share the common goal of understanding plant function, they are distinct in having their own intellectual histories and epistemologies."
Authors: Baolei Li and Jiaqi Sun.
Plant Physiology (2024)
Excerpt: "The functions of DIVARICATA (DIV) proteins, which belong to the MYB-like TF family, remain elusive (Fang et al., 2018). Six DIV genes have been identified in Arabidopsis based on their structural resemblance to the tomato I-box binding factor SlMYBI (Machemer et al., 2011). Among these, only DIV2 has been characterized as a positive regulator of seed germination in response to ABA and salt stress (Fang et al., 2018). However, the roles of the other DIV genes in the process of seed germination in response to salinity stress remain largely unexplored. In this issue of Plant Physiology, Zhang et al. (2024) provide evidence that DIV1 functions as a crucial integrative regulator that links salinity stress with ABA/GA signaling during seed germination."
"In summary, Zhang et al. (2024) illustrate the significant role of DIV1 as a positive regulator of Arabidopsis seed germination in response to salinity stress. DIV1 exerts a direct influence on ABA/GA signaling by promoting GASA4 expression and suppressing DOGL3 expression. Additionally, it indirectly modulates the expression of a series of germination-associated genes.
Authors: Diego Alejandro Gutiérrez-Villamil, Stanislav Magnitskiy and Helber Enrique Balaguera-López.
Postharvest Biology and Technology (2024)
Highlights • BR have a potential signaling function in the regulation of fruit development. • BR are a postharvest technology to regulate the ripening of fruits and vegetables. • BR mitigate chilling injury by increasing antioxidant systems. • BR induce expression genes of defense and immunity against postharvest pathogens.
Abstract: "Fresh horticultural products satisfy the nutritional, and industrial needs of consumers worldwide. However, the lack of understanding of the fruit development process, the accelerated senescence process and the lack of post-harvest technology in some regions, present a threat to the food and economic security of the food agribusiness. Brassinosteroids (BR) are plant hormones involved in the regulation of various physiological processes and have recently proven to be a viable post-harvest technology alternative to regulate the ripening and senescence of fruits and vegetables. In this review, the current state of BR research on fruit growth and development, physicochemical changes during ripening, and biotic-abiotic stress during the post-harvest life of horticultural products is presented. Furthermore, the review encompasses the effect of the application of exogenous BR and its relationship with molecular signaling on the processes mentioned above, including aspects such as methods, moments and BR analogues at the time of application, and the molecular mechanisms involved. This review proposes a basis for research of the physiological aspects of BR regulation in fruits and vegetables during their development and post-harvest period, and also points to a direction for in-depth investigation of the molecular mechanisms."
Authors: Faisal Zulfiqar, Anam Moosa, Hayssam M. Ali, Núria F. Bermejo and Sergi Munné-Bosch.
Plant Physiology and Biochemistry (2024)
Highlights: • Biostimulants are biobased solutions to tackle modern agriculture problems • For tackling climate change issue, biostimulants are not enough • Stopping wars and conflicts is the main solution to save environment and agriculture
Abstract: "Climate change is currently one of the main concerns of the agricultural sector, as it limits crop production and quality. Furthermore, the current context of global crisis with international political instability and war conflicts over the world is pushing the agricultural sector even more to urgently boost productivity and yield and doing so in a sustainable way in the current frame of climate change. Biostimulants can be an effective tool in alleviating the negative effects of environmental stresses to which plants are exposed, such as drought, salinity, heavy metals and extreme temperatures. Biostimulants act through multiple mechanisms, modifying gene expression, metabolism and phytohormone production, promoting the accumulation of compatible solutes and antioxidants and mitigating oxidative stress. However, it is important to keep in mind that the use and effect of biostimulants has limitations and must be accompanied by other techniques to ensure crop yield and quality in the current frame of climate change, such as proper crop management and the use of other sustainable resources. Here, we will not only highlight the potential use of biostimulants to face future agricultural challenges, but also take a critical look at their limitations, underlining the importance of a broad vision of sustainable agriculture in the current context of climate change."
Authors: Rui Yan, Tianle Zhang, Yuan Wang, Wenxiu Wang, Rahat Sharif, Jiale Liu, Qinglong Dong, Haoan Luan, Xuemei Zhang, Han Li, Suping Guo, Guohui Qi and Peng Jia.
Plant Physiology and Biochemistry (2024)
Highlights: • Seventeen GA2-oxidase genes identified in apple clustered into four clades. • MdGA2ox7 responded to cold and salt treatments. • MdGA2ox7 was activated during light-induced anthocyanin accumulation. • MdGA2ox7 alleviated cold and salt stress damage. • MdGA2ox7 promoted anthocyanin biosynthesis.
Abstract: "Apple (Malus domestica Borkh.) is a widely cultivated fruit crop worldwide but often suffers from abiotic stresses such as salt and cold. Gibberellic acid (GA) plays a pivotal in controlling plant development, environmental adaptability, and secondary metabolism. The GA2-oxidase (GA2ox) is responsible for the deactivation of bioactive GA. In this study, seventeen GA2-oxidase genes were identified in the apple genome, and these members could be clustered into four clades based on phylogenetic relationships and conserved domain structures. MdGA2ox7 exhibited robust expression across various tissues, responded to cold and salt treatments, and was triggered in apple fruit peels via light-induced anthocyanin accumulation. Subcellular localization prediction and experiments confirmed that MdGA2ox7 was located in the cytoplasm. Overexpression of MdGA2ox7 in Arabidopsis caused a lower level of active GA and led to GA-deficient phenotypes, such as dwarfism and delayed flowering. MdGA2ox7 alleviated cold and salt stress damage in both Arabidopsis and apple in concert with melatonin (MT). Additionally, MdGA2ox7 enhanced anthocyanin biosynthesis in apple calli and activated genes involved in anthocyanin synthesis. These findings provide new insights into the functions of apple GA2ox in regulating development, stress tolerance, and secondary metabolism."
Authors: Barbora Ndreca, Alison Huttly, Sajida Bibi, Carlos Bayon, George Lund, Joshua Ham, Rocío Alarcón-Reverte, John Addy, Danuše Tarkowská, Stephen Pearce, Peter Hedden, Stephen G. Thomas and Andrew L. Phillips.
BMC Plant Biology (2024)
Abstract: Background - Semi-dwarfing alleles are used widely in cereals to confer improved lodging resistance and assimilate partitioning. The most widely deployed semi-dwarfing alleles in rice and barley encode the gibberellin (GA)-biosynthetic enzyme GA 20-OXIDASE2 (GA20OX2). The hexaploid wheat genome carries three homoeologous copies of GA20OX2, and because of functional redundancy, loss-of-function alleles of a single homoeologue would not be selected in wheat breeding programmes. Instead, approximately 70% of wheat cultivars carry gain-of-function mutations in REDUCED HEIGHT 1 (RHT1) genes that encode negative growth regulators and are degraded in response to GA. Semi-dwarf Rht-B1b or Rht-D1b alleles encode proteins that are insensitive to GA-mediated degradation. However, because RHT1 is expressed ubiquitously these alleles have pleiotropic effects that confer undesirable traits in some environments. Results - We have applied reverse genetics to combine loss-of-function alleles in all three homoeologues of wheat GA20OX2 and its paralogue GA20OX1 and evaluated their performance in three years of field trials. ga20ox1 mutants exhibited a mild height reduction (approximately 3%) suggesting GA20OX1 plays a minor role in stem elongation in wheat. ga20ox2 mutants have reduced GA1 content and are 12–32% shorter than their wild-type segregants, comparable to the effect of the Rht-D1b ‘Green Revolution’ allele. The ga20ox2 mutants showed no significant negative effects on yield components in the spring wheat variety ‘Cadenza’. Conclusions - Our study demonstrates that chemical mutagenesis can expand genetic variation in polyploid crops to uncover novel alleles despite the difficulty in identifying appropriate mutations for some target genes and the negative effects of background mutations. Field experiments demonstrate that mutations in GA20OX2 reduce height in wheat, but it will be necessary to evaluate the effect of these alleles in different genetic backgrounds and environments to determine their value in wheat breeding as alternative semi-dwarfing alleles.
Authors: Jorge Hernández-García, Antonio Serrano-Mislata, María Lozano-Quiles, Cristina Úrbez, María A. Nohales, Noel Blanco-Touriñán, Huadong Peng, Rodrigo Ledesma-Amaro and Miguel A. Blázquez.
PNAS (2024)
Significance: DELLA proteins are plant-specific transcriptional hubs integrating environmental signals with endogenous cues. In order to regulate downstream processes, DELLAs modulate the activity of hundreds of transcription factors (TFs) and transcriptional regulators in various ways. Here, we describe the molecular mechanism underlying DELLA coactivator function. We show that DELLAs act as transcriptional activators by interacting with the Mediator complex subunit MED15. This interaction is necessary to regulate a specific subset of DELLA-dependent responses that are mediated by transcriptional coactivation, but not those regulated by TF sequestration. We further show that this mechanism is present in bryophyte DELLAs and thus represents a conserved mechanism of DELLA function in land plants.
Abstract: "DELLA proteins are negative regulators of the gibberellin response pathway in angiosperms, acting as central hubs that interact with hundreds of transcription factors (TFs) and regulators to modulate their activities. While the mechanism of TF sequestration by DELLAs to prevent DNA binding to downstream targets has been extensively documented, the mechanism that allows them to act as coactivators remains to be understood. Here, we demonstrate that DELLAs directly recruit the Mediator complex to specific loci in Arabidopsis, facilitating transcription. This recruitment involves DELLA amino-terminal domain and the conserved MED15 KIX domain. Accordingly, partial loss of MED15 function mainly disrupted processes known to rely on DELLA coactivation capacity, including cytokinin-dependent regulation of meristem function and skotomorphogenic response, gibberellin metabolism feedback, and flavonol production. We have also found that the single DELLA protein in the liverwort Marchantia polymorpha is capable of recruiting MpMED15 subunits, contributing to transcriptional coactivation. The conservation of Mediator-dependent transcriptional coactivation by DELLA between Arabidopsis and Marchantia implies that this mechanism is intrinsic to the emergence of DELLA in the last common ancestor of land plants."
Authors: Vojtěch Schmidt, Roman Skokan, Thomas Depaepe, Katarina Kurtović, Samuel Haluška, Stanislav Vosolsobě, Roberta Vaculíková, Anthony Pil, Petre Ivanov Dobrev, Václav Motyka, Dominique Van Der Straeten and Jan Petrášek.
Nature Communications (2024)
Editor's view: Here, the authors show that the biosynthesis of many compounds in green algae preceded their recruitment in phytohormone signaling and metabolism in land plants.
Abstract: "The genomes of charophyte green algae, close relatives of land plants, typically do not show signs of developmental regulation by phytohormones. However, scattered reports of endogenous phytohormone production in these organisms exist. We performed a comprehensive analysis of multiple phytohormones in Viridiplantae, focusing mainly on charophytes. We show that auxin, salicylic acid, ethylene and tRNA-derived cytokinins including cis-zeatin are found ubiquitously in Viridiplantae. By contrast, land plants but not green algae contain the trans-zeatin type cytokinins as well as auxin and cytokinin conjugates. Charophytes occasionally produce jasmonates and abscisic acid, whereas the latter is detected consistently in land plants. Several phytohormones are excreted into the culture medium, including auxin by charophytes and cytokinins and salicylic acid by Viridiplantae in general. We note that the conservation of phytohormone biosynthesis and signaling pathways known from angiosperms does not match the capacity for phytohormone biosynthesis in Viridiplantae. Our phylogenetically guided analysis of established algal cultures provides an important insight into phytohormone biosynthesis and metabolism across Streptophyta. Genomic evidence dates the origins of most phytohormones to terrestrialization or later."
Authors: Cong Li, Yanhang Chen, Qing Hu, Xiaolan Yang, Yunfeng Zhao, Yan Lin, Jianbo Yuan, Jinbao Gu, Yang Li, Jin He, Dong Wang, Bin Liu and Zhen-Yu Wang.
Plant Physiology (2024)
One-sentence summary: A pseudo-response regulator from the circadian system regulates a transcription factor involved in the abscisic acid-mediated drought stress response in soybean.
Abstract: "The circadian system plays a pivotal role in facilitating the ability of crop plants to respond and adapt to fluctuations in their immediate environment effectively. Despite the increasing comprehension of PSEUDO-RESPONSE REGULATORs (PRRs) and their involvement in the regulation of diverse biological processes, including circadian rhythms, photoperiodic control of flowering, and responses to abiotic stress, the transcriptional networks associated with these factors in soybean (Glycine max (L.) Merr.) remain incompletely characterized. In this study, we provide empirical evidence highlighting the significance of GmPRR3b as a crucial mediator in regulating the circadian clock, drought stress response, and abscisic acid (ABA) signaling pathway in soybeans. A comprehensive analysis of DNA affinity purification sequencing and transcriptome data identified 795 putative target genes directly regulated by GmPRR3b. Among them, a total of 570 exhibited a significant correlation with the response to drought, and eight genes were involved in both the biosynthesis and signaling pathways of ABA. Notably, GmPRR3b played a pivotal role in the negative regulation of the drought response in soybeans by suppressing the expression of abscisic acid responsive element-binding factor 3 (GmABF3). Additionally, the overexpression of GmABF3 exhibited an increased ability to tolerate drought conditions, and it also restored the hypersensitive phenotype of the GmPRR3b overexpressor. Consistently, studies on the manipulation of GmPRR3b gene expression and genome editing in plants revealed contrasting reactions to drought stress. The findings of our study collectively provide compelling evidence that emphasizes the significant contribution of the GmPRR3b-GmABF3 module in enhancing drought tolerance in soybean plants. Moreover, the transcriptional network of GmPRR3b provides valuable insights into the intricate interactions between this gene and the fundamental biological processes associated with plant adaptation to diverse environmental conditions."
Authors: Mahamud Hossain Al-Mamun, Christopher Ian Cazzonelli and Priti Krishna.
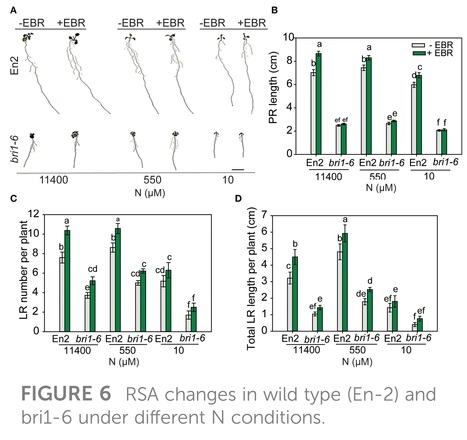
Frontiers in Plant Science (2024)
Abstract: "Plants modify their root system architecture (RSA) in response to nitrogen (N) deficiency. The plant steroidal hormone, brassinosteroid (BR), plays important roles in root growth and development. This study demonstrates that optimal levels of exogenous BR impact significant increases in lateral root length and numbers in Arabidopsis seedlings under mild N-deficient conditions as compared to untreated seedlings. The impact of BR on RSA was stronger under mild N deficiency than under N-sufficient conditions. The BR effects on RSA were mimicked in dominant mutants of BZR1 and BES1 (bzr1-1D and bes1-D) transcription factors, while the RSA was highly reduced in the BR-insensitive mutant bri1-6, confirming that BR signaling is essential for the development of RSA under both N-sufficient and N-deficient conditions. Exogenous BR and constitutive activity of BZR1 and BES1 in dominant mutants led to enhanced root meristem, meristematic cell number, and cortical cell length. Under mild N deficiency, bzr1-1D displayed higher fresh and dry shoot weights, chlorophyll content, and N levels in the shoot, as compared to the wild type. These results indicate that BR modulates RSA under both N-sufficient and N-deficient conditions via the transcription factors BES1/BZR1 module and confers tolerance to N deficiency."
|
Authors: Jingyi Han (撖静宜), Thomas Welch, Ute Voß, Teva Vernoux, Rahul Bhosale and Anthony Bishopp.
iScience (2024)
Highlights • The first intron of ARF7 is required for expression in the root apical meristem. • It is not required for expression in either lateral roots or the shoot apex. • We find no binding sites in the first intron required for expression in root tips. • We propose that this intron exerts its effects via Intron Mediated Enhancement.
Abstract: "Auxin regulates plant growth and development through the transcription factors of the AUXIN RESPONSE FACTOR (ARF) gene family. ARF7 is one of five activators that bind DNA and elicit downstream transcriptional responses. In roots, ARF7 regulates growth, gravitropism and redundantly with ARF19, lateral root organogenesis. In this study we analyzed ARF7 cis-regulation, using different non-coding sequences of the ARF7 locus to drive GFP. We show that constructs containing the first intron led to increased signal in the root tip. Although bioinformatics analyses predicted several transcription factor binding sites in the first intron, we were unable to significantly alter expression of GFP in the root by mutating these. We instead observed the intronic sequences needed to be present within the transcribed sequences to drive expression in the root meristem. These data support a mechanism by which Intron Mediated Enhancement regulates the tissue specific expression of ARF7 in the root meristem."
Authors: Byron Rusnak, Frances K. Clark, Batthula Vijaya Lakshmi Vadde and Adrienne H.K. Roeder.
Annual Review of Cell and Developmental Biology (2024)
Abstract: "One of the fundamental questions in developmental biology is how a cell is specified to differentiate as a specialized cell type. Traditionally, plant cell types were defined based on their function, location, morphology, and lineage. Currently, in the age of single-cell biology, researchers typically attempt to assign plant cells to cell types by clustering them based on their transcriptomes. However, because cells are dynamic entities that progress through the cell cycle and respond to signals, the transcriptome also reflects the state of the cell at a particular moment in time, raising questions about how to define a cell type. We suggest that these complexities and dynamics of cell states are of interest and further consider the roles signaling, stochasticity, cell cycle, and mechanical forces play in plant cell fate specification. Once established, cell identity must also be maintained. With the wealth of single-cell data coming out, the field is poised to elucidate both the complexity and dynamics of cell states."
Author: Daniel J. Cosgrove.
Annual Review of Plant Cell and Development (2024)
Abstract: "Expansins comprise an ancient group of cell wall proteins ubiquitous in land plants and their algal ancestors. During cell growth, they facilitate passive yielding of the wall's cellulose networks to turgor-generated tensile stresses, without evidence of enzymatic activity. Expansins are also implicated in fruit softening and other developmental processes and in adaptive responses to environmental stresses and pathogens. The major expansin families in plants include α-expansins (EXPAs), which act on cellulose-cellulose junctions, and β-expansins, which can act on xylans. EXPAs mediate acid growth, which contributes to wall enlargement by auxin and other growth agents. The genomes of diverse microbes, including many plant pathogens, also encode expansins designated expansin-like X. Expansins are proposed to disrupt noncovalent bonding between laterally aligned polysaccharides (notably cellulose), facilitating wall loosening for a variety of biological roles."
Authors: Feimei Guo, Minghui Lv, Jingjie Zhang and Jia Li.
Plant and Cell Physiology (2024)
Abstract: "Brassinosteroids (BRs) are a group of polyhydroxylated phytosterols that play essential roles in regulating plant growth and development as well as stress adaptation. It is worth noting that BRs do not function alone, but rather they crosstalk with other endogenous signaling molecules, including the phytohormones auxin, cytokinins (CKs), gibberellins (GAs), abscisic acid (ABA), ethylene (ET), jasmonates (JAs), salicylic acid (SA), and strigolactones (SLs), forming elaborate signaling networks to modulate plant growth and development. BRs interact with other phytohormones mainly by regulating each others’ homeostasis, transport, or signaling pathway at the transcriptional and posttranslational levels. In this review, we focus our attention on current research progress in BR signal transduction and the crosstalk between BRs and other phytohormones."
Authors: Da Zhang, Tan He, Xumin Wang, Chenchen Zhou, Youpeng Chen, Xin Wang, Shixiang Wang, Shuangcheng He, Yuan Guo, Zijin Liu and Mingxun Chen.
Plant Physiology (2024)
One-sentence summary: A MYB-like transcription factor positively modulates seed germination in response to salinity stress by regulating the expression of germination-associated genes.
Abstract: "Seed germination is a critical checkpoint for plant growth under unfavorable environmental conditions. In Arabidopsis (Arabidopsis thaliana), the abscisic acid (ABA) and gibberellic acid (GA) signaling pathways play important roles in modulating seed germination. However, the molecular links between salinity stress and ABA/GA signaling are not well understood. Herein, we showed that the expression of DIVARICATA1 (DIV1), which encodes a MYB-like transcription factor, was induced by GA and repressed by ABA, salinity, and osmotic stress in germinating seeds. DIV1 positively regulated seed germination in response to salinity stress by directly regulating the expression of DELAY OF GERMINATION 1-LIKE 3 (DOGL3) and GA-STIMULATED ARABIDOPSIS 4 (GASA4) and indirectly regulating the expression of several germination-associated genes. Moreover, NUCLEAR FACTOR-YC9 (NF-YC9) directly repressed the expression of DIV1 in germinating seeds in response to salinity stress. These results help reveal the function of the NF-YC9–DIV1 module and provide insights into the regulation of ABA and GA signaling in response to salinity stress during seed germination in Arabidopsis."
Authors: Davide Marzi, Patrizia Brunetti, Shashank Sagar Saini, Gitanjali Yadav, Giuseppe Diego Puglia and Raffaele Dello Ioio.
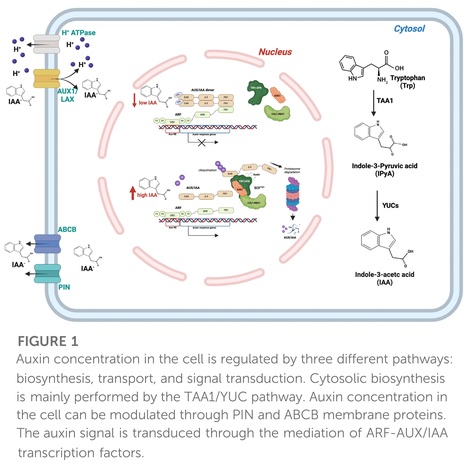
Frontiers in Genetics (2024)
Abstract: "Global climate change (GCC) is posing a serious threat to organisms, particularly plants, which are sessile. Drought, salinity, and the accumulation of heavy metals alter soil composition and have detrimental effects on crops and wild plants. The hormone auxin plays a pivotal role in the response to stress conditions through the fine regulation of plant growth. Hence, rapid, tight, and coordinated regulation of its concentration is achieved by auxin modulation at multiple levels. Beyond the structural enzymes involved in auxin biosynthesis, transport, and signal transduction, transcription factors (TFs) can finely and rapidly drive auxin response in specific tissues. Auxin Response Factors (ARFs) such as the ARF4, 7, 8, 19 and many other TF families, such as WRKY and MADS, have been identified to play a role in modulating various auxin-mediated responses in recent times. Here, we review the most relevant and recent literature on TFs associated with the regulation of the biosynthetic, transport, and signalling auxin pathways and miRNA-related feedback loops in response to major abiotic stresses. Knowledge of the specific role of TFs may be of utmost importance in counteracting the effects of GCC on future agriculture and may pave the way for increased plant resilience."
Authors: Kumi Matsuura-Tokita, Takamasa Suzuki, Yusuke Kimata, Yumiko Takebayashi, Minako Ueda, Takeshi Nakano, Hitoshi Sakakibara, Akihiko Nakano, and Tetsuya Higashiyama.
bioRxiv (2024)
Abstract: "Brassinosteroids (BRs) are steroid hormones identified in plants. Besides promoting cell elongation and division, BRs facilitate the development of both male and female reproductive tissues. In animals, reproductive steroid hormones play an essential role in reproductive tissue development by regulating gene expression. Here, we focused on the function of BRs during fertilization. We measured the content of biologically active BRs, brassinolide (BL) and castasterone (CS), in the reproductive tissues of Arabidopsis thaliana. Both BL and CS accumulated abundantly in pollen grains and in larger amounts in pistils than in leaves. To evaluate BL function during fertilization, we used an in vitro guidance assay with exogenously applied BL. Although pollen tubes need to be elongated through the pistils for efficient capacitation, BL treatment promoted pollen tube capacitation and improved attraction to ovules in vitro. Transcriptome analysis demonstrated that BL treatment induced the expression of half of the genes expressed in pollen tubes that elongated through the pistils. These results indicated that BL supplied from pistils is a key factor for pollen tube capacitation. However, using the bri1 mutant for the guidance assay resulted in reduced pollen tube capacitation, suggesting that BRI1-signaling in pistils is also important. Furthermore, BRs act on ovules. Exogenous BL application to ovules maintained guidance capacity by promoting the expression of small secreted proteins involved in pollen tube attraction and gamete fusion. Overall, BRs play a significant role as male and female reproductive hormones throughout the plant fertilization process."
Authors: Sonia Gazzarrini and Liang Song.
Annual Review of Plant Biology (2024)
Abstract: "Development is a chain reaction in which one event leads to another until the completion of a life cycle. Phase transitions are milestone events in the cycle of life. LEAFY COTYLEDON1 (LEC1), ABA INSENSITIVE3 (ABI3), FUSCA3 (FUS3), and LEC2 proteins, collectively known as LAFL, are master transcription factors (TFs) regulating seed and other developmental processes. Since the initial characterization of the LAFL genes (44, 71, 98–100), more than three decades of active research has generated tremendous amounts of knowledge about these TFs, whose roles in seed development and germination have been comprehensively reviewed (63, 43, 65, 81). Recent advances in cell biology and genetic and genomic tools have allowed the characterization of the LAFL regulatory networks in previously challenging tissues at a higher throughput and resolution in reference species and crops. In this review, we provide a holistic perspective by integrating advances at the epigenetic, transcriptional, posttranscriptional, and protein levels to exemplify the spatiotemporal regulation of the LAFL networks in Arabidopsis seed development and phase transitions, and we briefly discuss the evolution of these TF networks."
Authors: Wei Liu, Yuyan Cheng and Ziqiang Zhu (2024)
Journal of Plant Growth Regulation (2024)
Abstract: "The role of auxin in plant root thermomorphogenesis is still under debate. Here, we showed that auxin is necessary for root elongation in either seedling roots or detached roots. Our study clarified the uncertainty in the field and shed new light on future research in plant response to high ambient temperature."
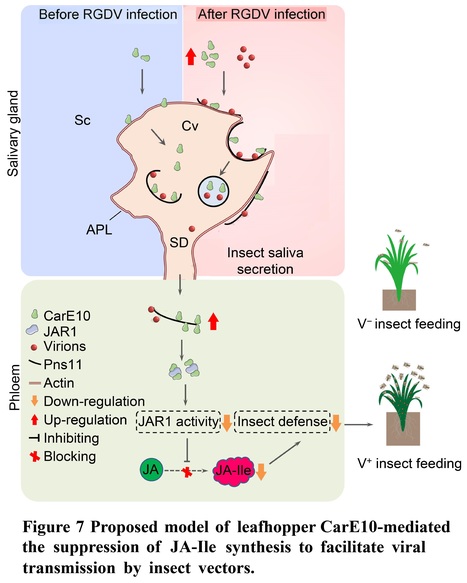
Authors: Yunhua Chi, Hongxiang Zhang, Siyu Chen, Yu Cheng, Xiaofeng Zhang, Dongsheng Jia, Qian Chen, Hongyan Chen and Taiyun Wei.
Plant Communications (2024)
Abstract: "Plant jasmonoyl-L-isoleucine (JA-Ile) is a major defense signal against insect feeding, but whether or how insect salivary effectors suppress JA-Ile synthesis and thus facilitate viral transmission in plant phloem remains elusive. Insect carboxylesterases (CarEs) are the third major family of detoxification enzymes. Here, we identify a new leafhopper CarE10 that specifically expressed in salivary glands and is secreted into rice phloem as the saliva component. Leafhopper CarE10 directly binds and promotes rice Jasmonate resistant 1 (JAR1) degradation by the proteasome system. Moreover, the direct association of CarE10 with JAR1 obviously impairs JAR1 enzyme activity for JA conversion to JA-Ile in in-vitro JA-Ile synthesis system. A devastating rice reovirus activates and promotes co-secretion of virions and CarE10 by virus-induced vesicles into saliva-stored salivary cavities of leafhopper vectors and ultimately into rice phloem to establish initial infection. Furthermore, virus-mediated increase of CarE10 secretion or overexpression of CarE10 in transgenic rice plants causes the reduced levels of JAR1 and thus suppresses JA-Ile synthesis, thereby promoting host attractiveness to insect vectors and facilitating initial viral transmission. Our findings provide insights into how insect salivary protein CarE10 suppresses host JA-Ile synthesis to benefit initial virus transmission in rice phloem."
Authors: Bihai Shi, Amelia Felipo-Benavent, Guillaume Cerutti, Carlos Galvan-Ampudia, Lucas Jilli, Geraldine Brunoud, Jérome Mutterer, Elody Vallet, Lali Sakvarelidze-Achard, Jean-Michel Davière, Alejandro Navarro-Galiano, Ankit Walia, Shani Lazary, Jonathan Legrand, Roy Weinstain, Alexander M. Jones, Salomé Prat, Patrick Achard and Teva Vernoux.
Nature Communications (2024)
Editor's view: Engineering of a biosensor allows the authors to map the signaling activity of the phytohormones gibberellins (GAs) and to show that GAs orient cell division at the shoot apex to establish the organization in parallel cell files of plant stems.
Abstract: "Growth at the shoot apical meristem (SAM) is essential for shoot architecture construction. The phytohormones gibberellins (GA) play a pivotal role in coordinating plant growth, but their role in the SAM remains mostly unknown. Here, we developed a ratiometric GA signaling biosensor by engineering one of the DELLA proteins, to suppress its master regulatory function in GA transcriptional responses while preserving its degradation upon GA sensing. We demonstrate that this degradation-based biosensor accurately reports on cellular changes in GA levels and perception during development. We used this biosensor to map GA signaling activity in the SAM. We show that high GA signaling is found primarily in cells located between organ primordia that are the precursors of internodes. By gain- and loss-of-function approaches, we further demonstrate that GAs regulate cell division plane orientation to establish the typical cellular organization of internodes, thus contributing to internode specification in the SAM."
Authors: Amanpreet Kaur, Norman B. Best, Thomas Hartwig, Josh Budka, Rajdeep Khangura, Steven McKenzie, Alejandro Aragón-Raygoza, Josh Strable, Burkhard Schulz and Brian P. Dilkes.
Plant Physiology (2023)
One-sentence summary: Molecular identity of a maize semidwarf mutant reveals a role for the maize GRAS domain transcription factor ortholog of DWARF AND LOW TILLERING in brassinosteroid signaling.
Abstract: "Brassinosteroids (BR) and gibberellins (GA) regulate plant height and leaf angle in maize (Zea mays). Mutants with defects in BR or GA biosynthesis or signaling identify components of these pathways and enhance our knowledge about plant growth and development. In this study, we characterized three recessive mutant alleles of GRAS transcription factor 42 (gras42) in maize, a GRAS transcription factor gene orthologous to the DWARF AND LOW TILLERING (DLT) gene of rice (Oryza sativa). These maize mutants exhibited semi-dwarf stature, shorter and wider leaves, and more upright leaf angle. Transcriptome analysis revealed a role for GRAS42 as a determinant of BR signaling. Analysis of the expression consequences from loss of GRAS42 in the gras42-mu1021149 mutant indicated a weak loss of BR signaling in the mutant, consistent with its previously demonstrated role in BR signaling in rice. Loss of BR signaling was also evident by the enhancement of weak BR biosynthetic mutant alleles in double mutants of nana plant1-1 and gras42-mu1021149. The gras42-mu1021149 mutant had little effect on GA-regulated gene expression, suggesting that GRAS42 is not a regulator of core GA signaling genes in maize. Single cell expression data identified gras42 expressed among cells in the G2/M phase of the cell cycle consistent with its previously demonstrated role in cell cycle gene expression in Arabidopsis (Arabidopsis thaliana). Cis-acting natural variation controlling GRAS42 transcript accumulation was identified by expression genome-wide association study (eGWAS) in maize. Our results demonstrate a conserved role for GRAS42/SCARECROW-LIKE 28 (SCL28)/DLT in BR signaling, clarify the role of this gene in GA signaling, and suggest mechanisms of tillering and leaf angle control by BR."
Authors: Kun Dong, Fuqing Wu, Siqi Cheng, Shuai Li, Feng Zhang, Xinxin Xing, Xin Jin, Sheng Luo, Miao Feng, Rong Miao, Yanqi Chang, Shuang Zhang, Xiaoman You, Peiran Wang, Xin Zhang, Cailin Lei, Yulong Ren, Shanshan Zhu, Xiuping Guo, Chuanyin Wu, Dong-Lei Yang, Qibing Lin, Zhijun Cheng and Jianmin Wan.
Molecular Plant (2024)
Abstract: "Although both protein arginine methylation (PRMT) and jasmonate (JA) signaling are crucial for regulating plant development, the relationship between these processes in spikelet development control remains unclear. Here, we utilized CRISPR/Cas9 technology to generate two OsPRMT6a loss-of-function mutants exhibiting various abnormal spikelet structures. Additionally, we found that OsPRMT6a could methylate arginine residues in the JA signal repressors OsJAZ1 and OsJAZ7. Arginine methylation of OsJAZ1 increased the affinity of OsJAZ1 for the JA receptors OsCOI1a and OsCOI1b in the presence of jasmonates (JAs), subsequently promoting the ubiquitination of OsJAZ1 by the SCFOsCOI1a/OsCOI1b complex and degradation via the 26S proteasome. This process ultimately released OsMYC2, a core transcriptional regulator in the JA signaling pathway, to activate or repress JA-responsive genes, thereby maintaining normal plant (spikelet) development. However, in the osprmt6a-1 mutant, reduced arginine methylation of OsJAZ1 impaired the interaction between OsJAZ1 and OsCOI1a/OsCOI1b in the presence of JAs. As a result, OsJAZ1 proteins became more stable, repressing JA responses, thus causing the formation of abnormal spikelet structures. Moreover, we discovered that JA signaling reduced the OsPRMT6a mRNA level in an OsMYC2-dependent manner, thereby establishing a negative feedback loop to balance JA signaling. Furthermore, we found that OsPRMT6a-mediated arginine methylation of OsJAZ1 likely serves as a switch to tune JA signaling to maintain normal spikelet development under harsh environmental conditions such as high temperatures. Thus, our study established a direct molecular link between arginine methylation and the JA signaling pathway.
|




 Your new post is loading...
Your new post is loading...


















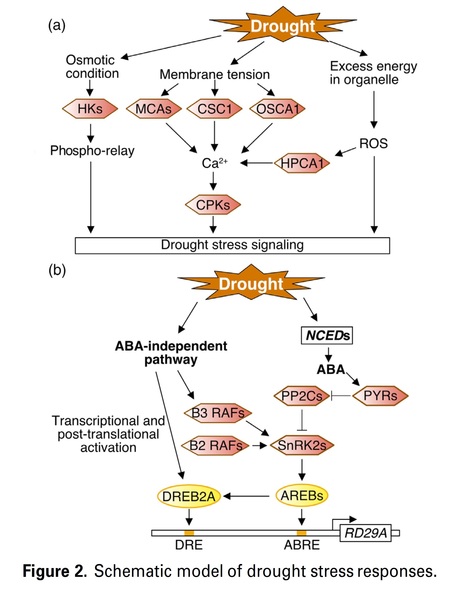
Authors: Hikaru Sato, Junya Mizoi, Kazuo Shinozaki and Kazuko Yamaguchi-Shinozaki.(2024)